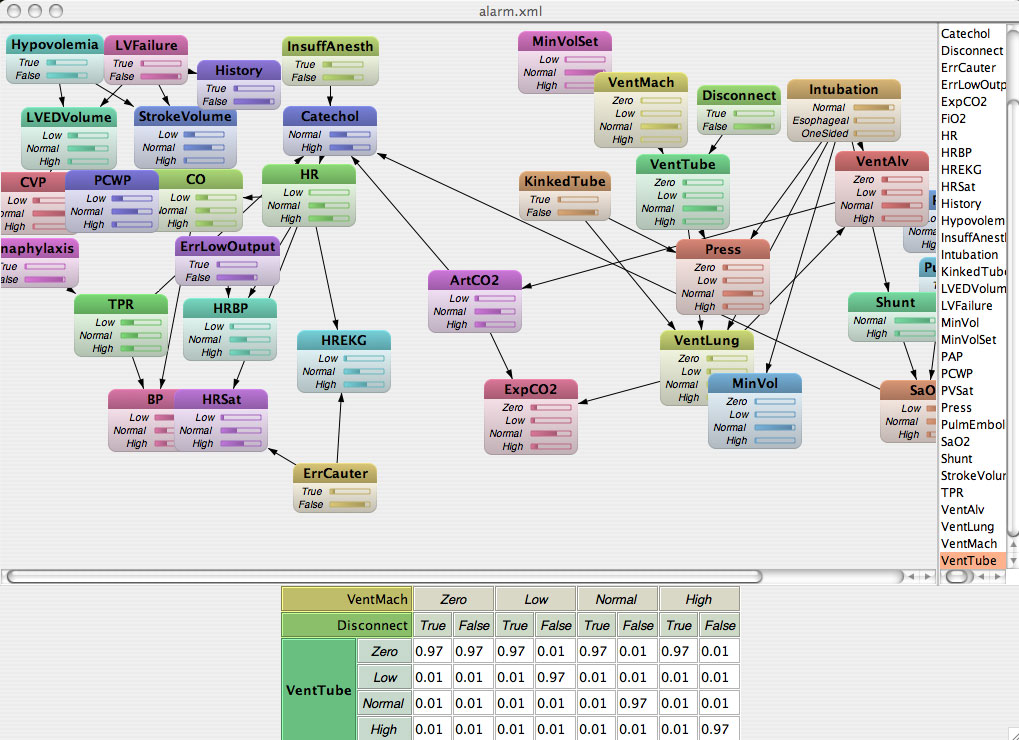
Inference in bayesian networks : first (good) results !
After trying to understand for hours why I didn’t get the good functions out of my junction tree inference algorithm implementation, I found a paper which gave me the missing key to this puzzle : the right way to decide in which clique’s potential to put each variable’s conditional probability function.
The key part was to choose a destination clique that contained all of the conditional probabilities function’s arguments (that is, all of the parents of the variable’s node in the graph). This is always possible because there is always such a clique, thanks to the moralization step (the moralization links all the parents of each node together, hence making them to be automatically in one or more cliques).
Download bayesian network inference engine prototype (Java 1.5 required)
The speed of computations is pretty good, especially since I switched the cliques potentials from composited functions (with a full algebraic tree embedded) to tabulated functions (with probabilities values available straight ahead in a table).
I dare not trust the values displayed yet but they look pretty accurate. For nodes without any parent there seems to be a few subtle differences between their values as defined in their probabilities tables and their inferred values. However, differences are very very small (about a tenth of a percent). It might come from small rounding errors during the inference process or from a bug (looks unlikely). I still need to compare results with other bayesian network engines, but for now, it just looks… perfect 😀